Prints effective sample sizes (ESS) and Geweke diagnostics for the scalar variance parameters in a fitted DBN model. Always run this before interpreting results to verify the MCMC sampler has converged.
Arguments
- results
Output from
dbn()
Details
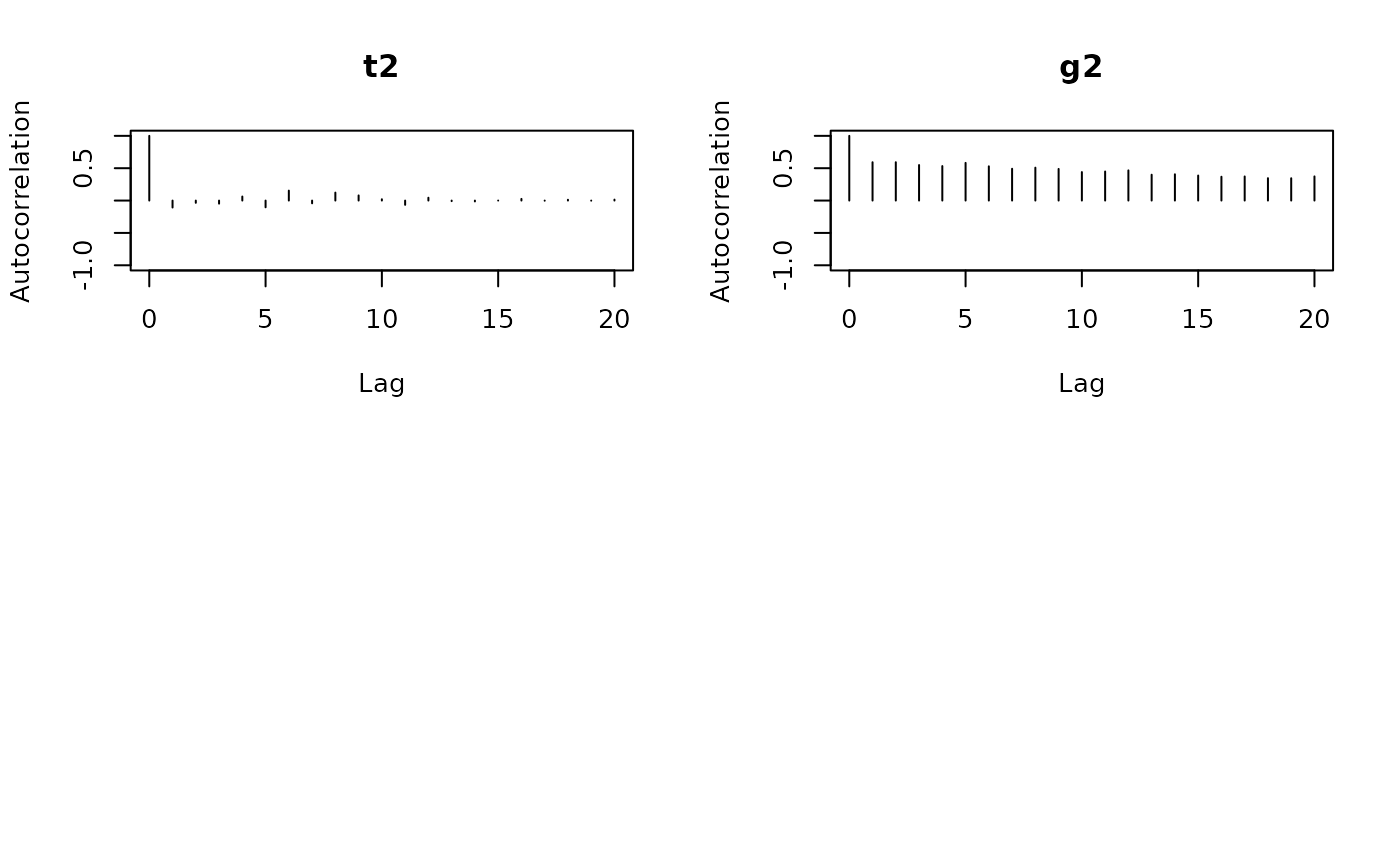
Effective sample size (ESS): The number of effectively independent
posterior draws. Values above 200-400 are generally adequate. Low ESS
means the chain is highly autocorrelated and you should increase
nscan or odens.
Geweke diagnostic: Tests whether the first and last portions of
the chain come from the same distribution. Absolute values above 2
suggest the chain has not converged and you should increase burn.
For visual diagnostics, use plot_trace() to inspect trace plots.
Examples
# \donttest{
sim <- simulate_static_dbn(n = 6, time = 10, seed = 1)
fit <- dbn(sim$Y, model = "static", nscan = 200, burn = 100, verbose = FALSE)
check_convergence(fit)
#> ℹ Fixed parameters (not sampled): "s2"
#>
#> ── Effective Sample Sizes
#> t2 g2
#> 246.975262 7.237814
#>
#> ── Geweke Diagnostic
#>
#> Fraction in 1st window = 0.1
#> Fraction in 2nd window = 0.5
#>
#> t2 g2
#> -2.115 -5.600
#>
# }