Regime-switching (HMM) bilinear network models
Tosin Salau & Shahryar Minhas
2026-03-28
Source:vignettes/hmm_dbn.Rmd
hmm_dbn.Rmd1 Overview
The HMM variant of the dynamic bilinear network model allows for regime-switching dynamics. At each time point, the network is governed by one of discrete regimes, each with its own influence matrices and . Transitions between regimes follow a Markov chain with an estimated transition matrix :
This model is appropriate when network dynamics exhibit abrupt
structural breaks and the analyst wants the model to discover when those
breaks occur, rather than specifying them in advance. For known break
points, the piecewise model provides more stable inference with fewer
parameters (see vignette("piecewise_dbn")).
Regime identification improves with network size, time series length, and the magnitude of the structural difference between regimes. For small networks () or subtle regime changes, the piecewise model with known break points is a more reliable choice.
2 Simulate a regime-switching network
We simulate a two-regime network with 12 actors over 30 time periods.
The high transition_prob (0.9) means each regime persists
for several consecutive periods before switching. The large
tau_A2 and tau_B2 values (0.8) create regimes
with substantially different dynamics, which is important for reliable
regime identification.
sim = simulate_hmm_dbn(
n = 12,
p = 1,
time = 30,
R = 2,
sigma2 = 0.3,
tau_A2 = 0.8,
tau_B2 = 0.8,
transition_prob = 0.9,
seed = 6886
)
dim(sim$Y)
#> [1] 12 12 1 30
# true regime sequence and frequencies
sim$S
#> [1] 2 2 2 2 2 2 2 2 2 1 2 2 2 2 2 2 2 2 2 1 1 1 1 1 1 1 1 1 1 1
table(sim$S)
#>
#> 1 2
#> 12 18The true regime labels show which regime generated each time point. With two regimes and a transition probability of 0.9, we expect each regime to persist for an average of 10 consecutive periods before switching.
3 Fit the HMM model
We set model = "hmm" and specify R = 2
regimes. The sampler jointly estimates regime-specific dynamics matrices
(,
),
the transition matrix
,
and the regime labels
.
fit = dbn(
sim$Y,
model = "hmm",
family = "ordinal",
R = 2,
nscan = 1000,
burn = 500,
odens = 2,
verbose = FALSE
)4 Model summary and transition matrix
The summary reports the posterior mean transition matrix and variance parameters. The diagonal of indicates regime persistence: values near 1 mean regimes tend to stick, while large off-diagonal values indicate frequent switching.
summary(fit)
#> Regime-switching (HMM) DBN model
#> nodes : 12
#> relations : 1
#> time pts : 30
#> regimes : 2
#>
#> mean 2.5% 97.5%
#> sigma2 1.000 1.000 1.000
#> tau_A2 0.041 0.037 0.046
#> tau_B2 0.039 0.035 0.043
#> g2 0.030 0.025 0.034
#>
#> Posterior mean transition matrix Pi:
#> [,1] [,2]
#> [1,] 0.691 0.309
#> [2,] 0.504 0.496Two things to note about the estimated
.
First, the model’s regime labels may not match the simulation’s — this
is the standard label-switching issue in mixture models and does not
affect the substantive results. Second, with the moderate chain length
used here (nscan = 1000), the transition matrix has
substantial posterior uncertainty. Recovering the true persistence (0.9
on the diagonal) requires longer chains and larger networks; with
actors and 30 time points, the model has limited data per regime to pin
down
precisely. For substantive work, run at least nscan = 5000
with burn = 2000.
5 Regime probabilities
The posterior probability of each regime at each time point provides
the model’s classification of the time series into regimes.
Probabilities near 0 or 1 indicate confident classification; values near
0.5 indicate ambiguity. Regime identification depends on how different
the regimes are (controlled by tau_A2 and
tau_B2 in the simulation) and how much data is available
per time point (number of actors).
plot_regime_probs(fit)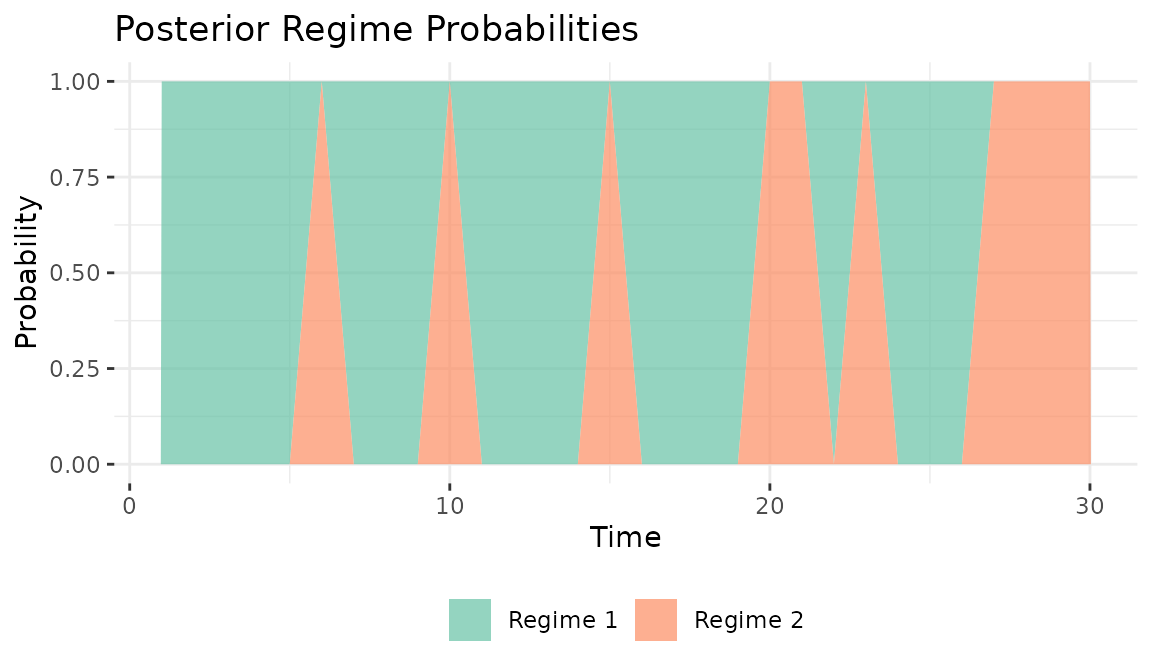
The regime_probs() function returns the underlying
probability matrix for custom analysis. Each row sums to 1 across
regimes.
probs = regime_probs(fit)
head(probs, 10)
#> Regime1 Regime2
#> Time1 1 0
#> Time2 1 0
#> Time3 1 0
#> Time4 1 0
#> Time5 1 0
#> Time6 0 1
#> Time7 1 0
#> Time8 1 0
#> Time9 1 0
#> Time10 0 16 Model diagnostics
The default plot() method for HMM fits provides a
combined diagnostic view: regime probabilities, the estimated transition
matrix, and MCMC trace plots for monitoring convergence.
plot(fit)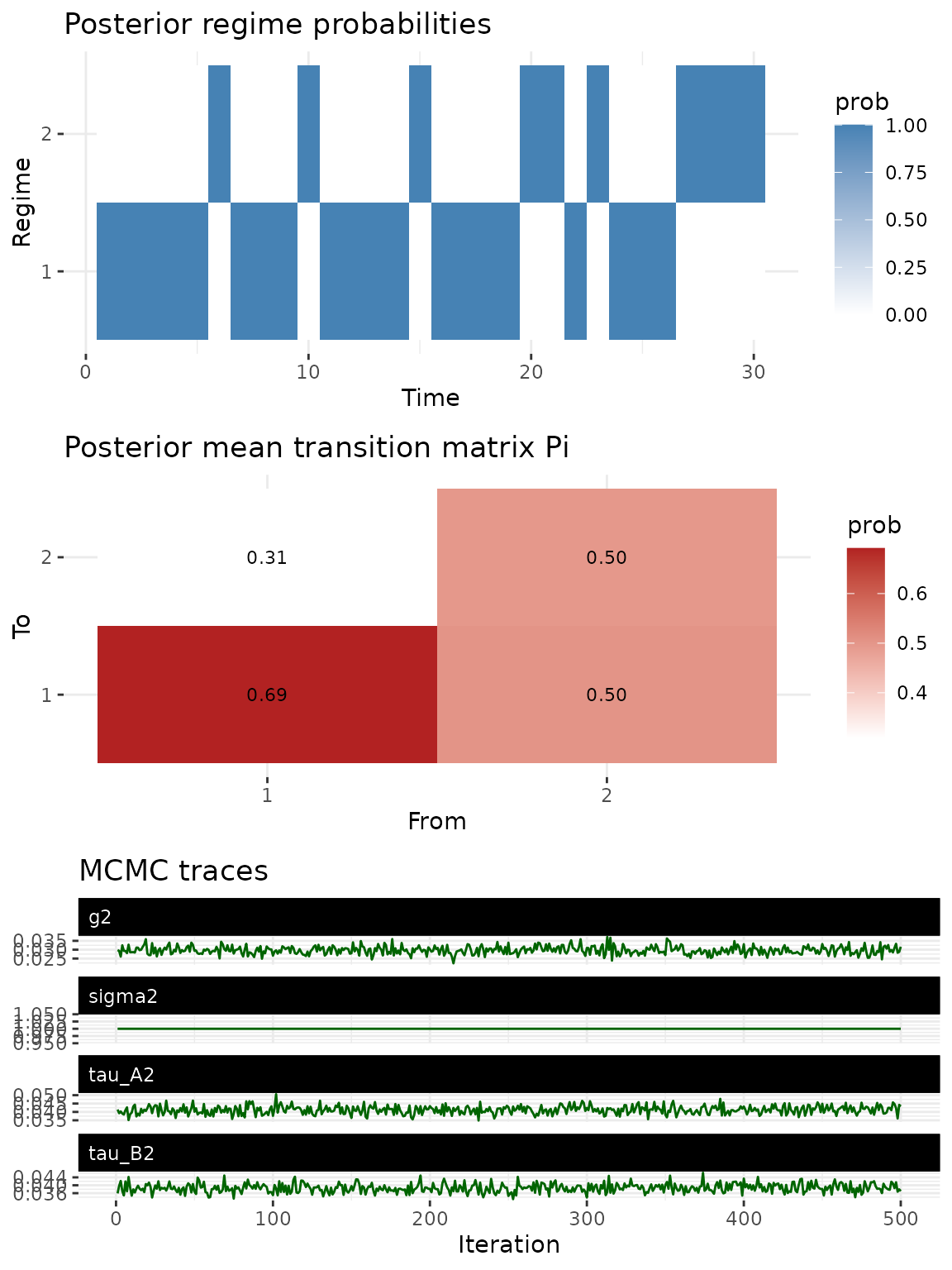
7 Dyad trajectory
The dyad_path() function tracks a specific bilateral
relationship over time with posterior credible intervals, averaging
across regime assignments.
dyad_path(fit, i = 2, j = 5)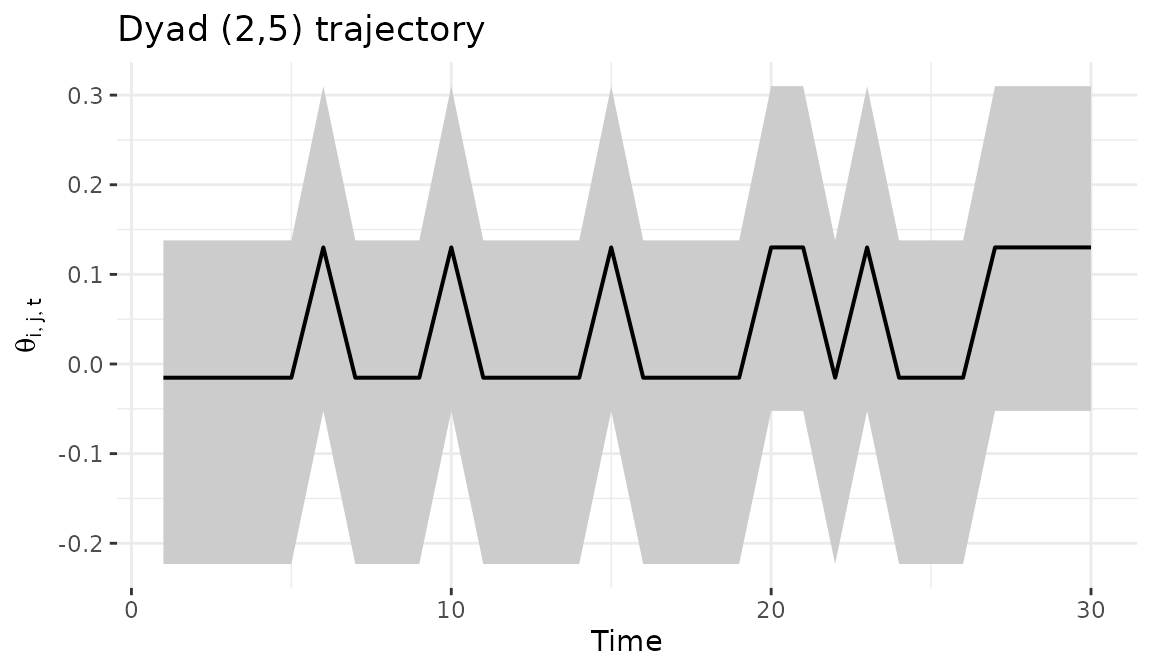
8 Forecasting
The predict() method samples future regime states from
and propagates the regime-specific bilinear dynamics forward. This means
forecast uncertainty reflects both posterior uncertainty in
and
and uncertainty about which regime will be active in future periods.
9 When to use HMM
The HMM model is appropriate when network dynamics exhibit discrete
structural breaks (such as policy changes, crises, or regime
transitions) and the analyst wants data-driven discovery of when those
breaks occur. The number of regimes
should be small (2-5), and the network should have enough actors and
time points to identify regime structure. For smoothly evolving dynamics
without discrete breaks, model = "dynamic" is a better
choice. For known break points, model = "piecewise"
provides more stable inference. For large networks (50+ actors),
model = "lowrank" offers better scalability.